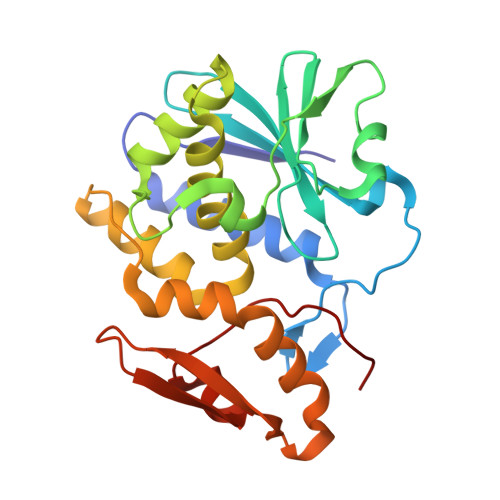
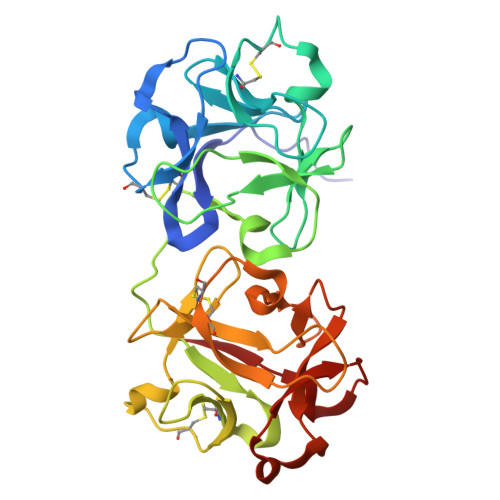
The Structure of the Type II Ribosome Inactivating Protein from Winter Aconite Eranthis hyemalis.
McConnell, M.-T., Talbert, R.C., Lee, J.C., Lisgarten, D.R., Lisgarten, J., Bertolo, E., Harvey, S.C., Levy, C.W., Lowe, E.D., Cooper, J.B., Naylor, C., Quiroz, R., Borges, R., Prince, S.M., Palmer, R.A.To be published.