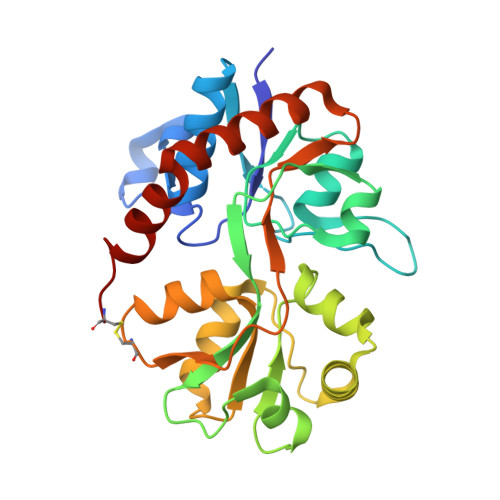
Three-Dimensional Structure of the Ligand-Binding Core of GluR2 in Complex with the Agonist (S)-ATPA: Implications for Receptor Subunit Selectivity.
Lunn, M.L., Hogner, A., Stensbol, T.B., Gouaux, E., Egebjerg, J., Kastrup, J.S.(2003) J Med Chem 46: 872-875
- PubMed: 12593667
- DOI: https://doi.org/10.1021/jm021020+
- Primary Citation of Related Structures:
1NNK, 1NNP - PubMed Abstract:
Two X-ray structures of the GluR2 ligand-binding core in complex with (S)-2-amino-3-(5-tert-butyl-3-hydroxy-4-isoxazolyl)propionic acid ((S)-ATPA) have been determined with and without Zn(2+) ions. (S)-ATPA induces a domain closure of ca. 21 degrees compared to the apo form. The tert-butyl moiety of (S)-ATPA is buried in a partially hydrophobic pocket and forces the ligand into the glutamate-like binding mode. The structures provide new insight into the molecular basis of agonist selectivity between AMPA and kainate receptors.
- Department of Medicinal Chemistry, Pharmaceutical University of Denmark, Universitetsparken 2, DK-2100 Copenhagen, Denmark.
Organizational Affiliation: