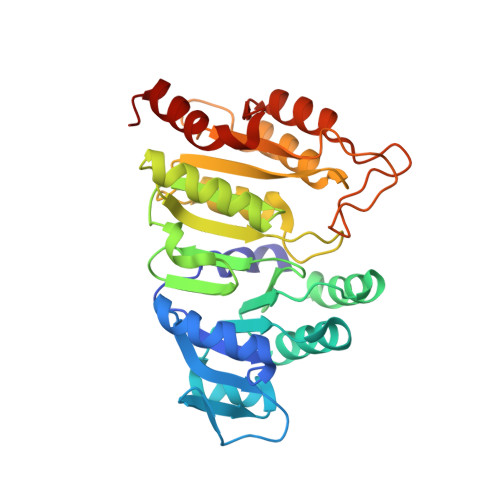
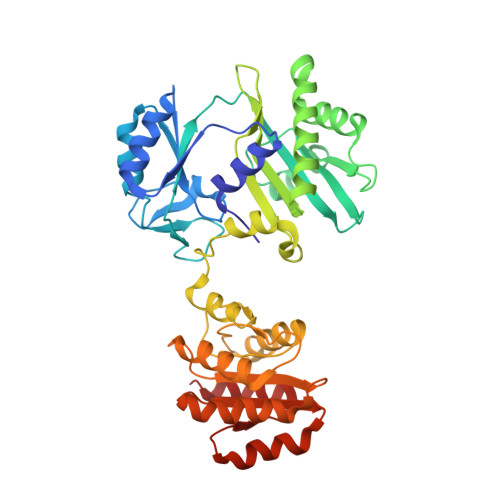
Interactions of GTP with the ATP-grasp Domain of GTP-specific Succinyl-CoA Synthetase
Fraser, M.E., Hayakawa, K., Hume, M.S., Ryan, D.G., Brownie, E.R.(2006) J Biological Chem 281: 11058-11065
- PubMed: 16481318
- DOI: https://doi.org/10.1074/jbc.M511785200
- Primary Citation of Related Structures:
2FP4, 2FPG, 2FPI, 2FPP - PubMed Abstract:
Two isoforms of succinyl-CoA synthetase exist in mammals, one specific for ATP and the other for GTP. The GTP-specific form of pig succinyl-CoA synthetase has been crystallized in the presence of GTP and the structure determined to 2.1 A resolution. GTP is bound in the ATP-grasp domain, where interactions of the guanine base with a glutamine residue (Gln-20beta) and with backbone atoms provide the specificity. The gamma-phosphate interacts with the side chain of an arginine residue (Arg-54beta) and with backbone amide nitrogen atoms, leading to tight interactions between the gamma-phosphate and the protein. This contrasts with the structures of ATP bound to other members of the family of ATP-grasp proteins where the gamma-phosphate is exposed, free to react with the other substrate. To test if GDP would interact with GTP-specific succinyl-CoA synthetase in the same way that ADP interacts with other members of the family of ATP-grasp proteins, the structure of GDP bound to GTP-specific succinyl-CoA synthetase was also determined. A comparison of the conformations of GTP and GDP shows that the bases adopt the same position but that changes in conformation of the ribose moieties and the alpha- and beta-phosphates allow the gamma-phosphate to interact with the arginine residue and amide nitrogen atoms in GTP, while the beta-phosphate interacts with these residues in GDP. The complex of GTP with succinyl-CoA synthetase shows that the enzyme is able to protect GTP from hydrolysis when the active-site histidine residue is not in position to be phosphorylated.
- Department of Biological Sciences, University of Calgary, Calgary, Alberta T2N 1N4, Canada. frasm@ucalgary.ca
Organizational Affiliation: