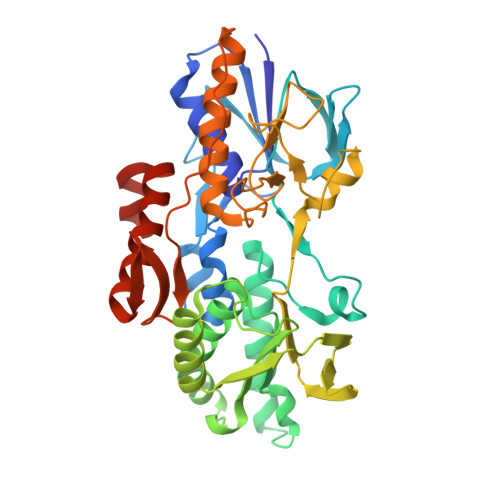
Crystal structure of type II NADH:quinone oxidoreductase from Caldalkalibacillus thermarum with an improved resolution of 2.15 angstrom.
Nakatani, Y., Jiao, W., Aragao, D., Shimaki, Y., Petri, J., Parker, E.J., Cook, G.M.(2017) Acta Crystallogr F Struct Biol Commun 73: 541-549
- PubMed: 28994401
- DOI: https://doi.org/10.1107/S2053230X17013073
- Primary Citation of Related Structures:
5WED - PubMed Abstract:
Type II NADH:quinone oxidoreductase (NDH-2) is a respiratory enzyme found in the electron-transport chain of many species, with the exception of mammals. It is a 40-70 kDa single-subunit monotopic membrane protein that catalyses the oxidation of NADH and the reduction of quinone molecules via the cofactor FAD. NDH-2 is a promising new target for drug development given its essential role in many bacterial species and intracellular parasites. Only two bacterial NDH-2 structures have been reported and these structures are at moderate resolution (2.3-2.5 Å). In this communication, a new crystallization platform is reported that produced high-quality NDH-2 crystals that diffracted to high resolution (2.15 Å). The high-resolution NDH-2 structure was used for in silico quinone substrate-docking studies to investigate the binding poses of menadione and ubiquinone molecules. These studies revealed that a very limited number of molecular interactions occur at the quinone-binding site of NDH-2. Given that the conformation of the active site is well defined, this high-resolution structure is potentially suitable for in silico inhibitor-compound screening and ligand-docking applications.
- Department of Microbiology and Immunology, University of Otago, 720 Cumberland Street, Dunedin 9054, New Zealand.
Organizational Affiliation: