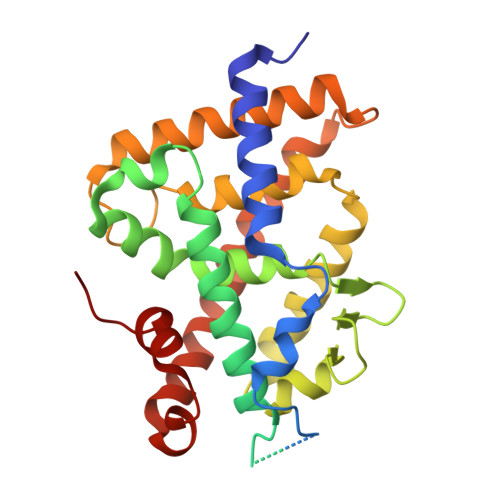
First Sila-Vitamin D Analogues: Design, Synthesis, Structural Analysis and Biological Activity.
Loureiro, J., Seoane, S., Sampaio-Dias, I.E., Peluso-Iltis, C., Guiberteau, T., Brito, B., Gregorio, C., Perez-Fernandez, R., Rochel, N., Mourino, A., Rodriguez-Borges, J.E.(2024) J Med Chem 67: 21505-21519
- PubMed: 39610329
- DOI: https://doi.org/10.1021/acs.jmedchem.4c02404
- Primary Citation of Related Structures:
9GY8, 9GYA, 9GYC, 9GYJ, 9GYK - PubMed Abstract:
The incorporation of silicon bioisosteres into pharmacological structures has been used as a strategy to improve the therapeutic potential of drugs. However, no secosteroidal silicon-containing VDR ligands have been developed. Here we report the design, synthesis, and biological activity of six analogues of the natural hormone 1,25-dihydroxyvitamin D3 (1,25D 3 ), which incorporate a silicon atom as a side chain-C25 isostere. The analogues were synthesized by the Wittig-Horner approach starting from Inhoffen-Lythgoe diol. The crystal structures of the complexes formed by the sila-analogues with the ligand binding domain of VDR revealed additional interactions of the sila-containing side chains that stabilize the VDR active conformation. These sila-analogues show similar VDR binding and transcriptional activity in comparison with the natural hormone 1,25D 3 , but with significantly less hypercalcemic activity. The new analogues, when combined with chemotherapy, significantly decrease cell proliferation.
- Department of Chemistry and Biochemistry, Faculty of Sciences, LAQV/REQUIMTE, University of Porto, Porto 4169-007, Portugal.
Organizational Affiliation: